Welcome to Scrimer¶
Scrimer is a GNU/Linux pipeline for designing PCR and genotyping primers from transcriptomic data.
Scrimer has been published in a peer-reviewed journal, please cite this if you use the pipeline or derive your own pipeline from our work:
Note
We present Scrimer as an end-to-end solution, from raw reads to usable primers. This is intended to help novice users, so they can better see the whole picture. However, this has an important downside - many steps in the pipeline are quite complex on their own (contig assembly, read mapping, variant calling etc.), and appropriate attention should be paid to checking the intermediate results.
In general, the most common solution for each given step is automatically chosen, using some reasonable default settings, but also giving the user the option to choose another program - using standard formats for input and output. The fine tuning of each step depending on the input data is up to the users.
Installation¶
Scrimer is a set of Python and Bash scripts that serve as a glue for several external programs.
The Python code is in the scrimer package, while Bash commands to run the Python scripts and the external
programs can be found in this documentation.
You can install Scrimer either to your own GNU/Linux machine, or use a prebuilt VirtualBox image:
Additionally a demo data set is available (it is already included in the VirtualBox image). It contains the whole project tree of one Scrimer run with all the intermediate files and an IGV session file, which can be used to load all the relevant tracks to IGV at once.
Pipeline workflow¶
The workflow is divided into several steps. The intermediate results of these steps can be visually inspected and checked for validity. There is no reason to continue with the process when a step fails, as further results will not be meaningful. The whole pipeline is a series of pre-made shell commands which are each supposed to be executed one after another by pasting them into the console.
The following pages describe these steps in detail, explain some choices we made and suggest ways how to check validity of the intermediate results.
Under normal circumsances it should take about a day to push your data through the pipeline, if everything goes well.
Scrimer dataflow¶
Dataflow diagram of the pipeline. Inputs are in green, processing steps in yellow and results in red. Arrows connecting steps are labelled with data format. This image was created using yEd.
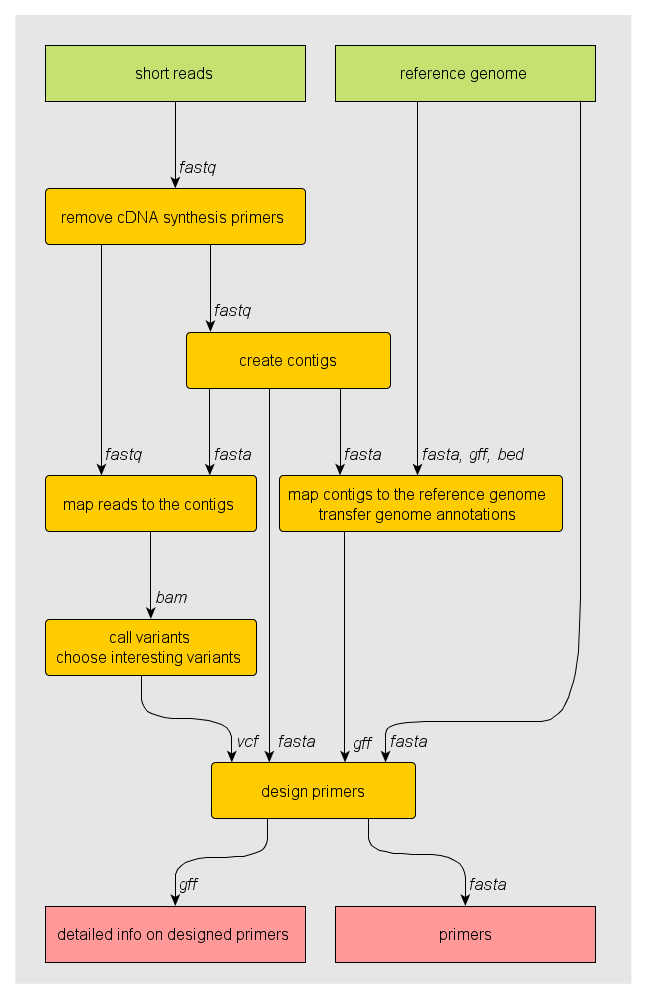
Scrimer components¶
Source code and reporting bugs¶
The source code and a bugtracker can be found at https://github.com/libor-m/scrimer.
License¶
Scrimer is licensed under the GNU Affero General Public License. Contact the author if you’re interested in other licensing terms.